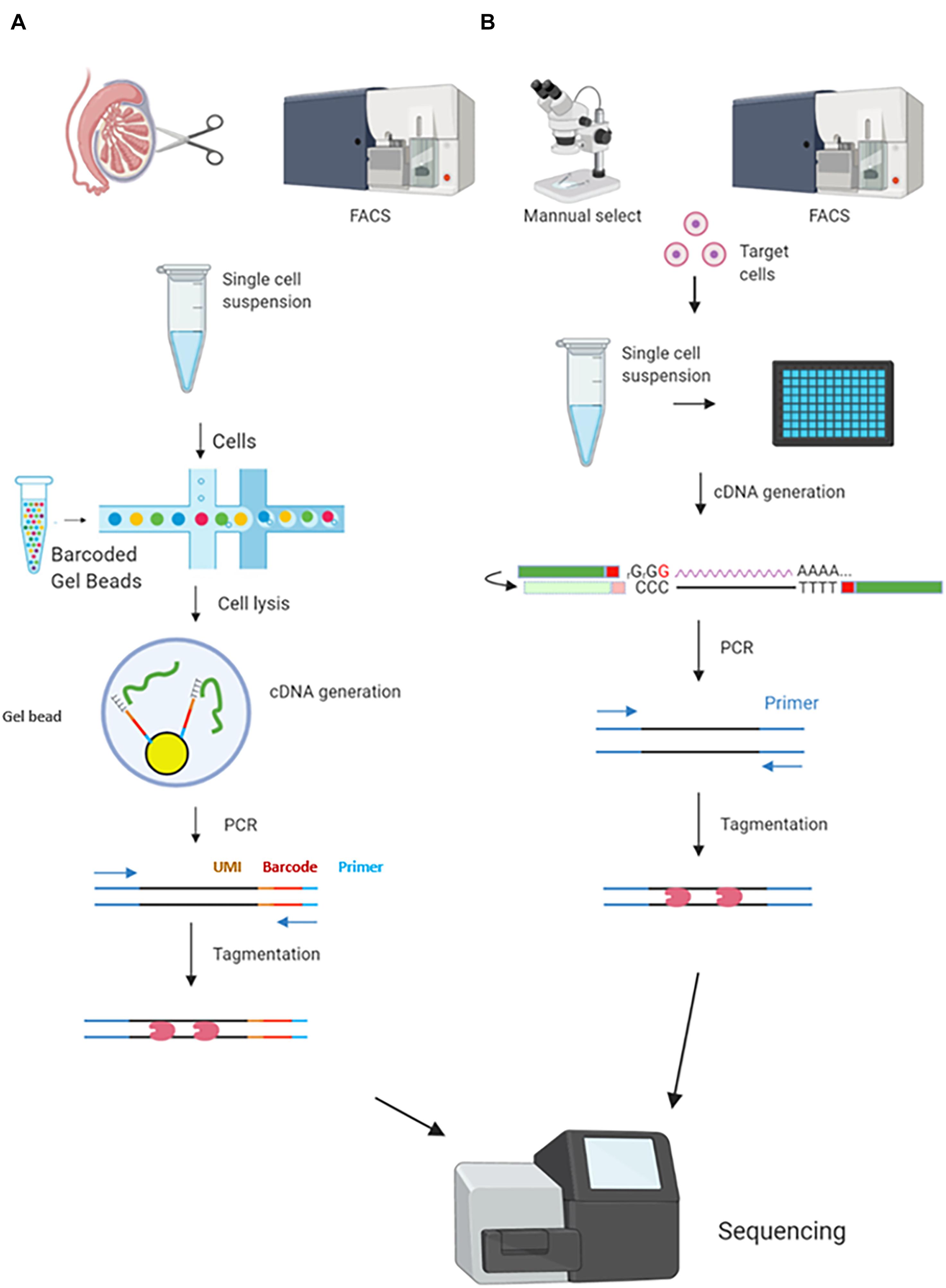
Single-Cell RNA Sequencing and Assay for Transposase-Accessible Chromatin Using Sequencing Reveals Cellular and Molecular Dynamics of Aortic Aging in Mice | Arteriosclerosis, Thrombosis, and Vascular Biology
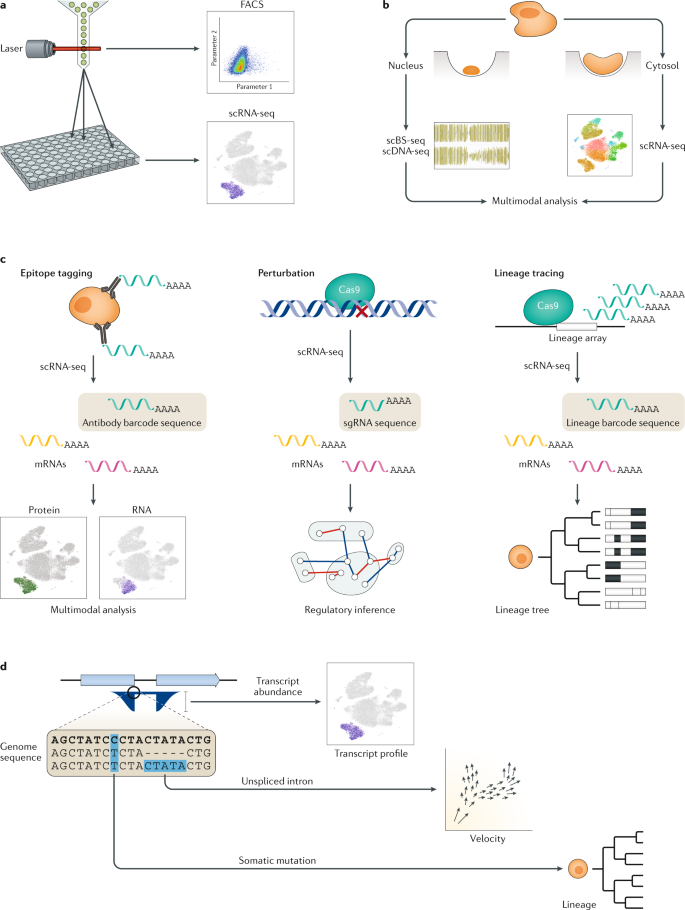
Single-cell RNA sequencing technologies and bioinformatics pipelines | Experimental & Molecular Medicine

Single-Cell RNA Sequencing to Disentangle the Blood System | Arteriosclerosis, Thrombosis, and Vascular Biology
IQCELL: A platform for predicting the effect of gene perturbations on developmental trajectories using single-cell RNA-seq data | PLOS Computational Biology

Using single-cell transcriptomics to understand functional states and interactions in microbial eukaryotes | Philosophical Transactions of the Royal Society B: Biological Sciences

Single-Cell Transcriptomics: A High-Resolution Avenue for Plant Functional Genomics: Trends in Plant Science

Single-cell technologies and analyses in hematopoiesis and hematological malignancies - Experimental Hematology

Using single‐cell genomics to understand developmental processes and cell fate decisions | Molecular Systems Biology

New avenues for systematically inferring cell-cell communication: through single-cell transcriptomics data | SpringerLink